|
6/20/2023 0 Comments Immune repertoire sequencing
Such assays permit a more precise understanding of the biological mechanisms leading to graft injury, and the opportunity for earlier clinical intervention, mitigating rejection and improving graft outcomes. We therefore require novel, less invasive and reliable assays for monitoring the alloimmune response in solid organ transplant to recognize immune events at the earliest possible time prior to the advent of irreversible organ injury. Unfortunately, current diagnostic methods are limited in utility and application, and typically only confirm rejection once significant organ damage has already occurred. Novel solid-phase or cellular assays detecting anti-human leukocyte antigen (HLA) antibodies help to discriminate between cellular (CMR) and antibody mediated rejection (AMR). Donor-derived cell-free DNA (dd-cfDNA) has shown potential as a biomarker for early detection of immune activation and serial surveillance ( 1). But by virtue of its invasive nature, biopsy is normally employed only when clinical data raise diagnostic suspicion. Transplant biopsy is the current gold standard for diagnosing rejection in solid organ transplant, combining histology with more detailed opportunities for cellular, molecular, and spatial biology evaluation of graft injury. 
Principal mechanisms of alloantigen recognition Mixed lymphocyte culture followed by TCR sequencing was used in 6 studies to define an alloreactive repertoire and in specialized transplant settings to track tolerance.Ĭonclusion: Methodological approaches to immune repertoire sequencing are becoming established and offer considerable potential as a novel clinical tool for pre- and post-transplant immune monitoring.ġ. Rejectors and those with opportunistic infections were more likely to have clonal expansion in T or B cell populations.  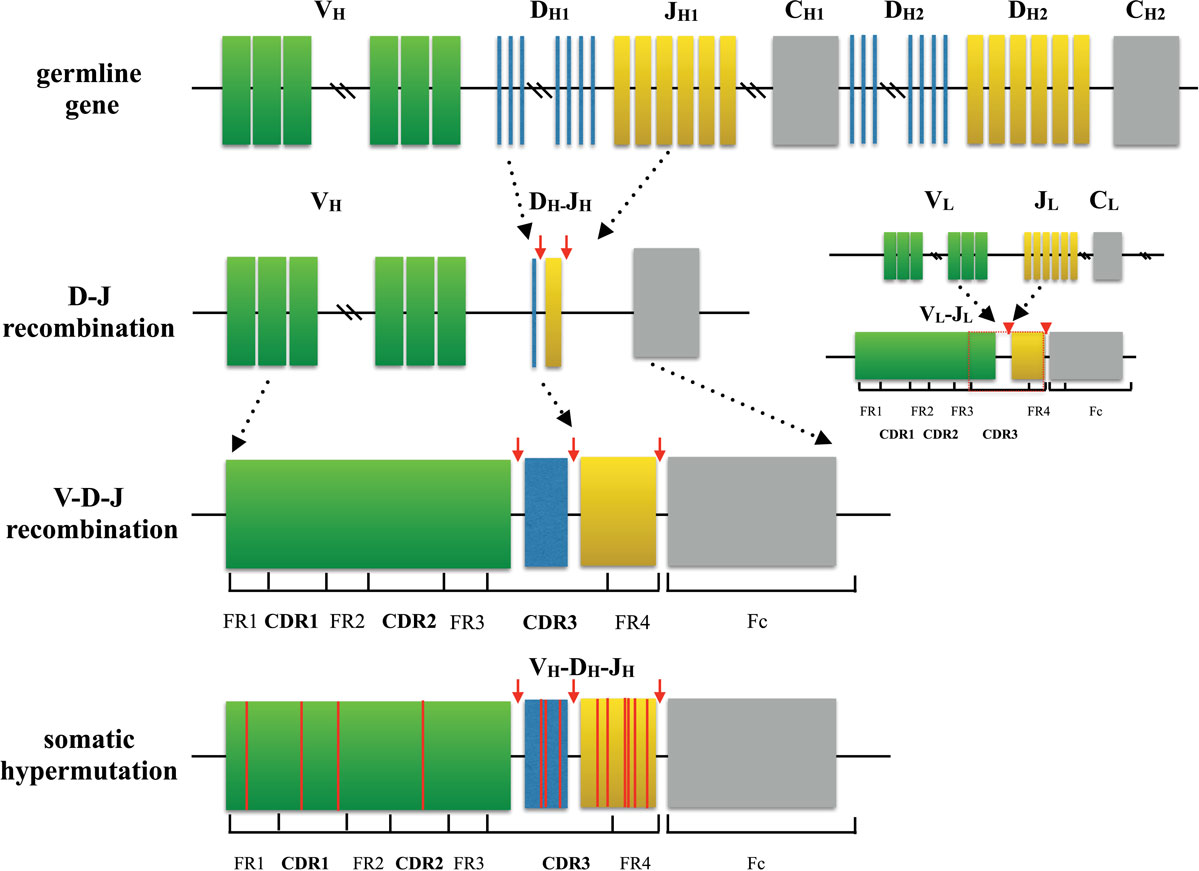
Repertoires of transplant recipients were found to have decreased diversity in both rejectors and non-rejectors when compared to healthy controls. The predominant method for repertoire characterization was sequencing the CDR3 region of the TCR β chain. Results: Our initial search yielded 1933 articles of which 37 met the inclusion criteria 16 of these were kidney transplant studies (43%) and 21 were other or general transplantation studies (57%). Data were extracted based on study and methodology characteristics. Manual filtering of the search results was performed based on relevancy and predefined inclusion criteria. Methods: We searched MEDLINE and PubMed Central for English-language studies published between 20 that examined T cell/B cell repertoire dynamics upon immune activation. Objective: We performed a review of current literature to examine research in immune repertoire sequencing in organ transplantation and to assess the feasibility of this technology for clinical application in immune monitoring. 6Department of Pediatrics, The Research Institute of the McGill University Health Centre and the Montreal Children’s Hospital, Montreal, QC, Canadaīackground: Measurement of T cell receptor (TCR) or B cell receptor (BCR) gene utilization may be valuable in monitoring the dynamic changes in donor-reactive clonal populations following transplantation and enabling adjustment in therapy to avoid the consequences of excess immune suppression or to prevent rejection with contingent graft damage and to indicate the development of tolerance.5Department of Epidemiology, Biostatistics and Occupational Health, McGill University, Montreal, QC, Canada.4Department of Medicine, Division of Nephrology, McGill University, Montreal, QC, Canada.3Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, BC, Canada.2Department of Urologic Sciences, University of British Columbia, Vancouver, BC, Canada.1Department of Medicine, University of British Columbia, Vancouver, BC, Canada.Sherwood 1,3, Franz Fenninger 1,3, Ruth Sapir-Pichhadze 4,5, Constantin Polychronakos 6, James Lan 1,3 and Paul A.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
